Center for Archaeological Science, Sichuan University published in top forensic science journal FSIG, reporting that global population non-CODIS STR profiles provide new ideas for forensic complex DNA analysis

Short tandem repeat (STR) accounts for about 6% of the genome and is a class of highly polymorphic genetic markers. Since its first application in forensic DNA analysis in the 1990s, STR has played an important role in population genetics research, personal identification, biogeographical ancestry inference, paternity testing, and kinship analysis. The current mainstream STR detection kits generally contain only 20-30 STR markers, including the core loci of the FBI Combined DNA Index System (CODIS), which are difficult to meet the increasingly complex needs of forensic DNA analysis. High-throughput next-generation sequencing (NGS) enables the joint detection of a large number of CODIS and non-CODIS STR genetic markers in a single experiment, thereby effectively improving the system effectiveness of the DNA analysis system and is expected to bring new breakthroughs to complex DNA analysis. However, due to the lack of a global population map of genetic variation in non-CODIS STRs, this technical route has not been widely used in actual cases.
Recently, the team of He Guanglin from the Institute of Rare Diseases, West China Hospital of Sichuan University and Center for Archaeological Science, Sichuan University, together with the team of Academician Liu Chao from the Guangdong Provincial Drug Experimental Technology Center of the Chinese Academy of Engineering, published a research paper entitled "Comprehensive landscape of non-CODIS STRs in global populations provides new insights into challenging DNA profiles" in the authoritative forensic journal Forensic Science International: Genetics, using 4150 high-depth whole genome sequencing (WGS) data from different populations around the world to comprehensively analyze the polymorphic characteristics of 178 non-CODIS STRs. On this basis, the study combined real and simulated data sets to systematically evaluate the application potential of large-scale non-CODIS STR systems in solving difficult forensic problems such as mixed DNA analysis of multiple contributors, biogeographical ancestry inference under mixed genetic backgrounds, and complex kinship identification. The study revealed for the first time the polymorphic pattern of global population genome non-COIDS STRs, proving that the inclusion of polymorphic non-CODIS STRs is of great significance for improving the ability of complex DNA analysis, and provides a scientific basis for the subsequent development of large-scale STR detection systems.
Research Results:
1. Construction of a whole-genome non-CODIS STR reference gene set
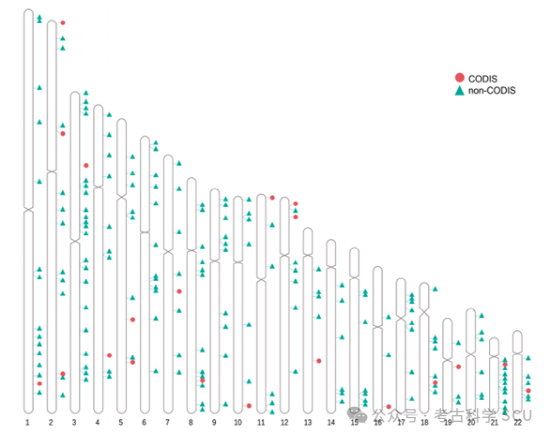
Figure 1. Chromosomal distribution of STR loci included in the study.
A total of 198 polymorphic STR loci were included in the study, including 20 CODIS core loci and 178 non-CODIS loci, and the genomic annotation information of all STRs was improved through sequence search, repetitive sequence alignment, and manual verification.
2. Evaluation of global population non-CODIS STR polymorphism and forensic application potential
Figure 2. Evaluation of CODIS and non-CODIS polymorphism and forensic parameters. a: Global population STR polymorphism; b: STR forensic application parameters. PIC, Polymorphism Information Content; He, Expected heterozygosity; Ho, Observed heterozygosity; MP, Match probability; PD, Power of discrimination; PE, Power of exclusion; TPI, Typical paternity index.
The study evaluated the polymorphism and forensic parameters of STR in different intercontinental populations. The results showed that non-CODIS STR has polymorphism and forensic efficacy comparable to CODIS STR. In addition, both CODIS and non-CODIS STR showed obvious population stratification: the African population (AFR) had higher overall polymorphism, while the Oceania population (OCN) had the lowest polymorphism.
3. Population structure and biogeographical ancestry inference
Considering the population polymorphism and linkage disequilibrium patterns, the study simulated a large-scale STR detection system (extdSTR) including 20 CODIS core loci and 88 non-CODIS loci, which realized the effective differentiation of the genetic background of different intercontinental populations around the world.

Figure 3. Global population genetic structure analysis based on 108 STRs. a: PCA analysis of five intercontinental populations; b: Non-metric multidimensional scaling analysis of 26 global populations based on the population differentiation index (Fst) matrix.
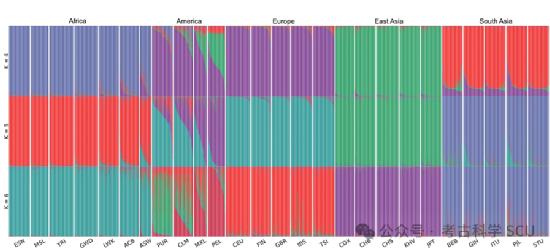
Figure 4. STRUCTURE unsupervised cluster analysis of 26 global populations (K=4, 5, 6).
Furthermore, the study constructed a random forest classification model based on 108 STRs to evaluate the biogeographical ancestry inference ability of the system. The results showed that the application of large-scale non-CODIS STR can effectively distinguish the biogeographical ancestry of individuals with different intercontinental ancestral backgrounds.
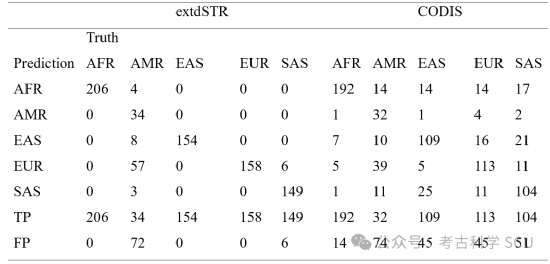
Table 1 Confusion matrix of the RF model for predicting ancestral origin based on extdSTR and CODIS core loci sets. TP: True Positive; FP: False Positive.
4. Mixed DNA analysis
In order to verify the ability of non-CODIS to analyze complex mixed DNA, the study simulated and generated a series of mixed DNA profiles composed of 2 to 8 contributors. The results showed that the inclusion of non-CODIS STR effectively improved the accuracy of contributor number (NoC) estimation and the evidence strength of mixed DNA profiles.
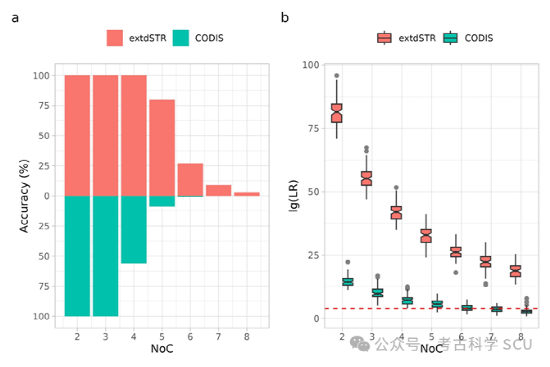
Figure 5. Deconvolution of DNA mixture profiles using extdSTR and CODIS core loci sets (n = 100). The true number of contributors NoC is 2-8. a: NoC inference accuracy evaluation; b: Likelihood ratio values of target individuals in mixed DNA profiles.
5. Complex kinship inference
Finally, the study evaluated the ability of non-CODIS to perform complex kinship inference. The results showed that large-scale non-CODIS can effectively distinguish between first- to third-degree relatives, but the ability to distinguish between relatives above the third degree is still poor.

This study systematically analyzed the polymorphism characteristics and forensic application potential of global population genome non-CODIS STRs using high-depth WGS data. The results showed that non-CODIS STRs have high polymorphism and forensic efficacy in different populations, and have broad application prospects in biogeographical ancestry inference, DNA mixture deconvolution, and complex kinship analysis. With the widespread application of NGS technology in the field of forensic medicine, incorporating non-CODIS STR into large-scale STR detection systems is expected to provide powerful tools for complex DNA analysis.
The first author of the paper is Huang Yugo, a postdoctoral fellow at the Institute of Rare Diseases, West China Hospital of Sichuan University, and the first corresponding author is He Guanglin, an associate researcher at the Institute of Rare Diseases, West China Hospital of Sichuan University and Center for Archaeological Science, Sichuan University. Academician Liu Chao of Guangdong Provincial Drug Experimental Technology Center (National Drug Laboratory Guangdong Branch) is the co-corresponding author.